X-ray crystallography sheds light on a protein crucial for chromosome segregation in bacteria
When cells divide, replicated chromosomes carrying DNA separate and move to daughter cells.
This process of chromosome segregation is essential in all domains of life to ensure each daughter cell inherits a full copy of the genetic information of the parent.
During segregation, DNA is either pulled or pushed to the daughter cells. All mechanisms discovered so far can be reduced to three core components: proteins that help organise and disentangle replicated chromosomes; a protein that consumes nucleotide triphosphate molecule to provide a directional force; and an adaptor protein that tethers the force-generating machinery to the DNA.
Adaptor protein has a crucial job in coordinating chromosome segregation and chromosome organisation. This task is not trivial, even in simpler organisms such as bacteria. In many bacteria, the three-component ParA-ParB-parS complex is responsible for chromosome segregation.
The parS DNA site is the first to be segregated following chromosome replication. DNA neighbouring to parS is occupied by hundreds of the adaptor protein called ParB, which then interacts with a protein partner, ParA, to drive chromosomes to daughter cells.
Despite the biological significance of this process, how multiple ParB can gather in the vicinity of parS is not yet fully understood.
To better understand how ParB works, researchers in the group of Dr Tung Le collaborated with the Protein Crystallography Platform at the John Innes Centre and the group of Dr Anjana Badrinarayanan at the National Centre for Biological Sciences (NCBS, Bangalore, India).
The team solved X-ray crystallography structures of a ParB molecule (from a freshwater bacterium Caulobacter crescentus) either bound to the parS DNA or bound to a small molecule called cytidine triphosphate (CTP).
The team showed that ParB is an open “clamp” when it first binds to parS DNA.
Upon binding to CTP, ParB then shuts a series of “molecular gates” to close the clamp, thereby wrapping itself around the DNA. The closed clamp of ParB can now slide away on the DNA, much like train on a track, to travel to the distal DNA region. Multiple ParB clamps can wrap around and move on DNA one after another, and this probably explains how hundreds of ParB gather by themselves near parS inside the cells.
The team also identified a ‘clamp-locked’ variant of ParB that once it is bound on DNA, cannot open the clamp to release the DNA. Interestingly, cells harbouring this ‘clamp-locked’ variant are unviable, suggesting that a dynamic cycle of closing and opening of the ParB clamp is essential for proper chromosome segregation.
Dr Tung Le said, “I am excited that we are a step closer to understanding how bacteria faithfully segregate their genetic information. The ParABS system is also essential for the transmission of many clinically relevant plasmids so the knowledge gained here might be useful to reduce the spread of plasmid-borne antibiotics resistance.
“It has also been a fantastic collaboration with Professor Dave Lawson and Dr Clare Stevenson at the X-ray crystallography platform at the John Innes Centre. Adam Jalal, the first author of this work, and I have learnt a lot from them. We are also incredibly grateful to Dr Anjana Badrinarayanan and Afroze Chimthanawala at the NCBS for performing some key experiments, especially when some techniques stopped working in our lab after the first COVID lockdown last year.”
- Read the paper; ‘A CTP-dependent gating mechanism enables ParB spreading on DNA‘, published in eLife
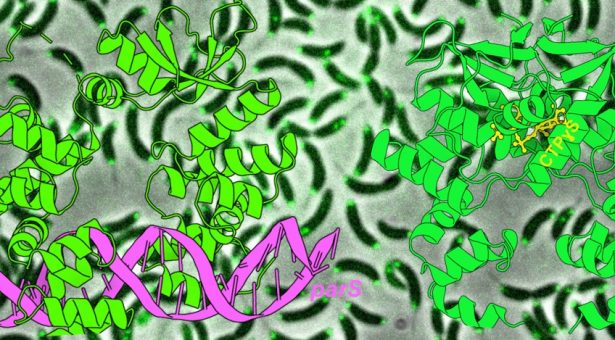
The image at the top of this page shows X-ray crystallography structures of ParB bound to DNA (magenta) or to CTPyS (yellow), and the location of the ParB inside Caulobacter crescentus cells.



