Engineered bacterium is helping in the search for new antibiotics
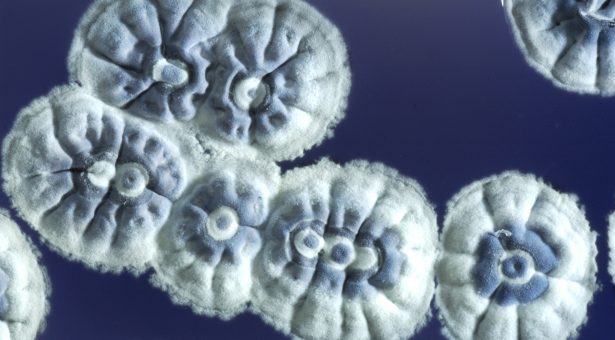
To help in the search for new antibiotics, Juan Pablo Gomez-Escribano and Professor Mervyn Bibb have adapted a strain of Streptomyces coelicolor to express clusters of genes found from genome sequencing projects whose functions are unknown.
The increasing problem of resistance to antibiotics in a broad range of human pathogens means that we are in growing need of new clinically effective antimicrobial agents.
One strategy to discover such compounds is ‘genome mining,’ the search for potential new antibiotic genes in sequenced genomes.
Sequencing the genomes of actinomycetes, the group of bacteria from which two thirds of current antibiotics are derived, has thrown up a large number of gene clusters of unknown function.
Some of these ‘cryptic’ gene clusters may produce antimicrobial compounds, but only under certain conditions, or in undetectable amounts in their native hosts.
As part of ActinoGen, an EU-funded project using genomics to exploit unexplored genetic resources in the search for new antibiotics, the John Innes Centre scientists have engineered Streptomyces coelicolor M145 to enhance the expression of these gene clusters and to characterise their products.
The strain has had its own antibiotic genes removed, to reduce competition for resources, and has also been engineered to increase the level of production from introduced antibiotic gene clusters. Both changes will make it much easier to identify and characterise the products of cryptic gene clusters.
The strains are now being used by a number of different research groups.
“We believe that these strains can make a major contribution to the discovery of novel natural products and may contribute significantly to addressing the urgent need for new antibiotics,” said Professor Bibb.