Lightweight tool boxes clever with heavyweight data

A free, open-source toolkit to help researchers deal with data management overload has been devised by the John Innes Centre Informatics team.
The toolkit, called ‘dtool’ is a suite of software for managing scientific data and making it accessible for front-line researchers working across multiple projects areas.
It works by packaging data and metadata – information that identifies the data – into handy boxes or datasets.
These self-contained packages of data and metadata make it easy to move datasets around and create remote backup copies.
The tools work with both traditional file systems as well as cloud options such as Amazon S3 and Microsoft Azure, allowing researchers to choose the storage solution that best suits their needs and budgets.
The system means researchers can quickly find datasets of interest without the headache of having to access and maintain a central database. The packaged metadata can be used to verify the integrity of the data in the box.
The John Innes Centre team outline the benefits of dtool in an article published in PeerJ – the Journal of Life and Environmental Sciences.
They say the toolkit offers peace of mind because researchers know data underpinning scientific results are safe, searchable and accessible across a highly distributed research environment such as the John Innes Centre.
Dr Tjelvar Olsson, Senior Scientific Data and Infrastructure Manager at the John Innes Centre, who is one of the creators of the system, said: “At the John Innes Centre we have 40 different research groups dealing with huge volumes of all kinds of data.
“We want more people to use dtool to manage their data. We have designed it in a way that slots into their way of working, a lightweight, solution used in a minimal kind of way that sits on top of what they are already doing.”
One of the early adopters of dtool is the team of Dr Brande Wulff working on disease resistance in wheat at the John Innes Centre.
Advances in technologies such as genetic sequencing and computational systems biology have contributed to an explosion in the volumes and types of data. While this has led to major advances in plant and microbial science it has led to substantial challenges in data management and processing for front line researchers.
Dr Matthew Hartley, Head of the JIC’s informatics team, who helped to devise dtool, said the impact was already being felt.
“Managing data at scale is one of the biggest challenges in computational biological research. dtool has made storing our data cheaper, given us peace of mind and sped up our research.”.
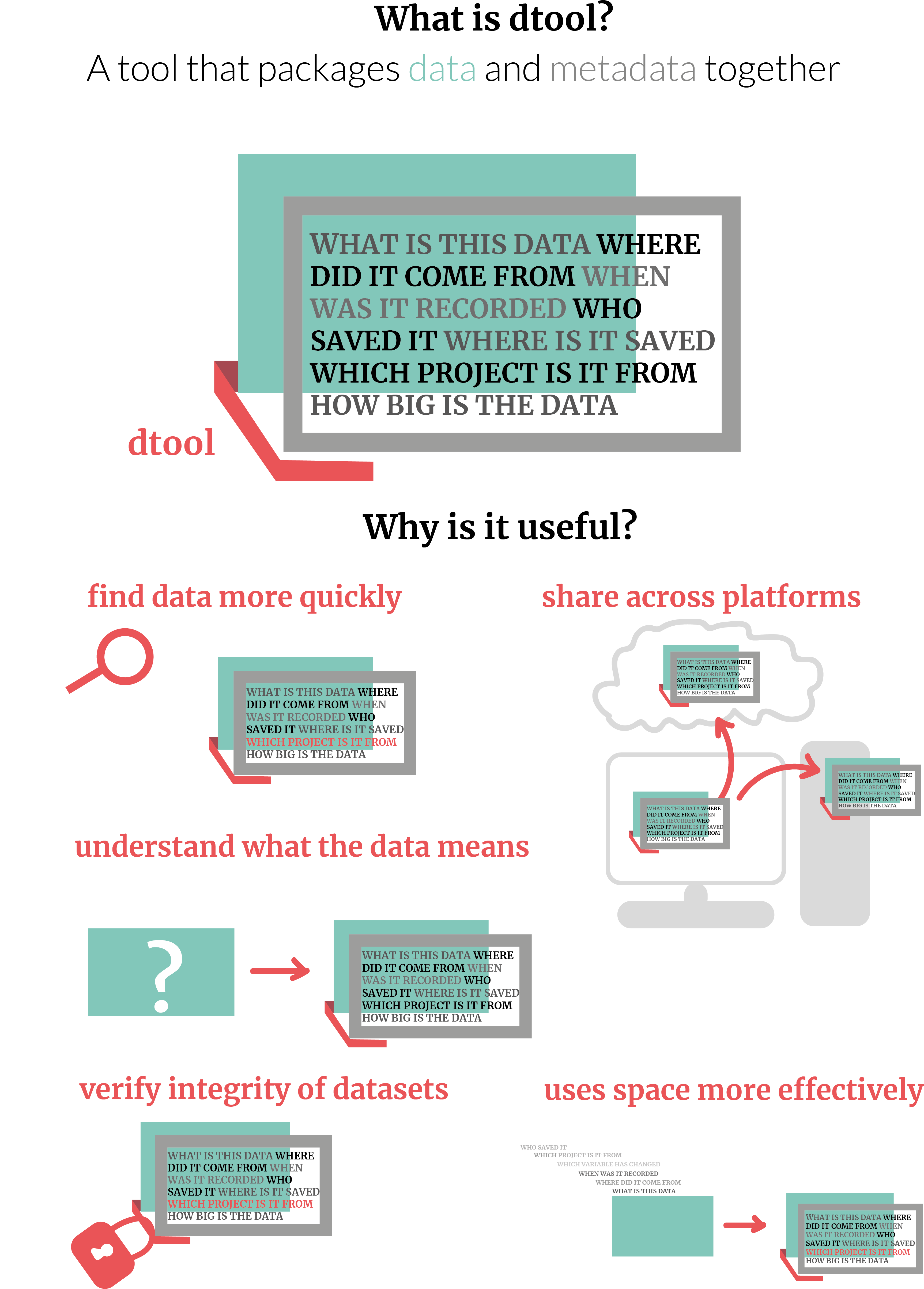
What is dtool?
Essentially dtool is a tool that packages data and metadata together.
Why is it useful?
- Find data more quickly
- Share across platforms
- Understand what the data means
- Verify the integrity of datasets
- Use space more effectively
