Meet the Molecules; Fasamycin and Formicamycin

Antimicrobial resistance poses a major threat to our society and we are fast approaching a situation reminiscent of the era before antibiotics.
In an attempt to retain the upper hand, scientists at the John Innes Centre and the UEA, are combining the latest technological advances with traditional antibiotic hunting techniques such as investigating interesting microbes to try and uncover new antibiotics.
New techniques for antibiotic hunting can be used to explore previously inaccessible bacterial habitats, to look for bacterial secondary metabolites. Sequencing technologies can then be used to uncover biosynthetic gene clusters (BCG) that may code for new clinically relevant antibiotics.
These techniques were applied to investigate ant nests in the thorny acacia trees in Kenya. Detailed exploration of these nests revealed a new species of bacteria called, Streptomyces formicae KY5, which was isolated from swabs of both the nests and ants themselves.
Back in the lab bioassay studies of Streptomyces formicae KY5 revealed several compounds which displayed potent activity against the Methicillin resistant pathogen Staphylococcus aureus, more commonly known as MRSA.
Encouragingly they were also effective against the Vancomycin resistant Enterococci (VRE) and other Gram-positive bacteria.
So far, three potential antibiotic compounds have been identified from Streptomyces formicae;
- 3 x Fasamycins (C-E)
- 10 x Formicamycins (A-J)
- 9 x Formicapyridines (A-I)
These three compounds are all produced from the same set of genes but differ in their chemistry to make them distinct from each other.
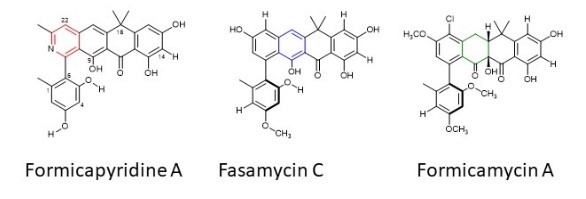
Promising results have been obtained from both fasamycin and formicamycin compounds which have both shown activity against MRSA and VRE. Currently it is unknown what role the formicapyridines play in bacterial interactions..
Furthermore, attempts to generate bacteria resistant to these compounds have so far proved difficult which may indicate a high barrier to resistance. This is of great importance as not only is the demand for new antibiotics increasing but antibiotics that are not easily made redundant by resistance may aid in solving long-term antimicrobial problems.
Current work in the Wilkinson lab is focussing on uncovering what organisms these compounds are active against and how these compounds work to inhibit pathogens.
This work is an example of working with bacteria to combat antimicrobial resistance with promising results.
The image at the top of this page is courtesy of Wikipedia Commons.