Super-sweet pea seeds
Starch is the main carbohydrate reserve in plants and a major source of nutrition for plants and animals.
Understanding starch biosynthesis and its genetics in pea was advancing rapidly in the late 1980s into the 1990s and the John Innes Centre was at the forefront of this research, with our scientists biochemically characterising two previously identified genes, which influence the starch content of the pea seed and, as part of a mutagenesis programme, identified three additional and novel starch mutants.
At the time of the mutagenesis programme, the genes at two loci that affect seed shape and starch content had already been identified.
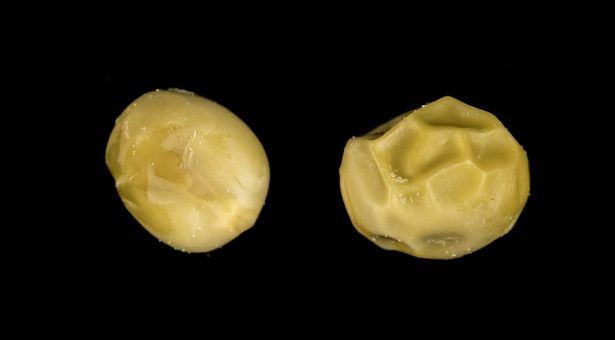
These were the r gene studied by Mendel and the rb gene. They both confer a wrinkled appearance to pea seeds and reduce the starch content of the seed due to the decreased activity of certain enzymes involved in starch synthesis (starch-branching enzyme, and ADP glucose pyrophosphorylase, respectively).
By screening induced mutant lines, Trevor Wang and Cliff Hedley were able to identify three new wrinkled-seeded mutant types – rug3, rug4, rug5 – and a round-seeded mutant, Iam, which are all affected in their starch content. Wang described how, much like Mendel, he spent years counting round and wrinkled seeds to separate the genetically wrinkled from the physiologically wrinkled to understand their genetics.
The rug3 mutant
The mutants at the rug3 locus were found to be the most severely affected in their starch content, lowering the seed values to almost negligible amounts depending on the allele involved.
The evidence from biochemical assays and linkage studies indicated that the pea mutant alleles decrease the activity of a phosphoglucomutase enzyme.
These rug3 mutants have extremely high sugar content, due to the severe reduction in starch synthesis, and yielded ‘super-sweet’ pea seeds that were also higher in vitamin C, which are highly desirable commercial traits.
Consequently, we collaborated with a frozen pea producer, who wanted to introduce the rug3 mutation into breeding lines to produce super-sweet frozen peas.
The rug3 gene had the added benefit of prolonging harvesting time, due to low or no starch synthesis. However, the low starch content also meant the seeds had few reserves to be drawn on when put under stress.
Ultimately, the producer did not develop the rug3 pea mutation commercially, and this breeding programme was eventually sold off. However, a pea cultivar carrying an independently derived rug3 mutation was successfully bred in Denmark. In 2016, this cultivar ‘Jedi’ has been trialled by the John Innes Centre and the yields from microplots were promising.
By the end of the 1990s, we had identified the major enzymes involved in starch synthesis in pea, aided by the identification of the range of mutant alleles.
The programme was successful as it developed an understanding of the enzymes and genetics involved in plant starch synthesis with potential for commercial use.
Further reading
- Harrison, C.J., Mould, R.M., Leech, M.J., Johnson, S.A., Turner, L., Schreck, S.L., Baird, K.M., Jack, P.L., Rawsthorne, S., Hedley, C.L., Wang, T.L. (2000). The rug3 locus of pea encodes plastidial phosphoglucomutase. Plant Physiology, 122, 1187-1192. DOI: org/ 10. 1104/ pp. 122. 4. 1187
- Wang, T.L., Hedley, C.L. (1991). Seed development in peas: knowing your three ‘r’s’ (or four, or five). Seed Science Research, 1, 3- 14. DOI: http://dx.doi.org/10.1017/S096025850000057X
- Wang, T.L., Bogracheva, T.Y., Hedley, C.L. (1998). Starch: as simple as A, B, C?. Journal of Experimental Botany, 49, 481-502. DOI: 10.1093/jxb/49.320.481
