The social networks of plant mitochondria
Alongside publishers Cambridge University Press we collaborate on the open access journal; Quantitative Plant Biology (QPB), which provides an interdisciplinary forum for high quality research on ground-breaking discoveries and predictions in plant science.
QPB is a dedicated home for research that applies techniques such as data mining and analysis, mathematical modelling and machine-learning to plant biology.
The journal welcomes research from across the spectrum of fundamental, applied and societal plant research; across all biological scales, from molecular through cellular and organismal to populations; and be based on data from laboratories, fieldwork and citizen science
With that in mind, this week we asked University of Birmingham PhD Student Joanna Chustecki to guest blog on her work using computational analysis to map the social networks of plant mitochondria.
We started by asking Joanna, why she became interested in mitochondria and what she means by their social networks.
“Mitochondria are amazing organelles. They are highly dynamic, and viewing their motion under the microscope is incredible- I clearly remember my first time watching them.
Where are they going? What are they doing talking to all those other organelles? How are they moving? We’re getting closer to the answers.
Our work looks at the population of mitochondria in plant cells as a whole and there is a lot of information to be gained by taking a broad, systemic approach.
Networks have previously been used to characterise all kinds of systems – cellular organisation in plants, the structure of the endoplasmic reticulum, fungal networks, and even reticulated mitochondrial networks in yeast.
But we are taking a different approach because plant mitochondria spend most of their time alone, as discrete individuals. But they do interact, coming into close contact with one another- we’re building encounter networks from these events, and using them to quantify connectivity across all of the individuals.
This given us an insight into not only when they communicate, but why.
We uncovered that the plant cell has a constant trade-off to surmount; how to keep mitochondria together for exchange of proteins, mtDNA, metabolites etc, and how to keep them apart to avoid local mutagen build up, get energy delivered around the cell quickly and meet up with other organelles.
We used modelling approaches and mutants to explore this relationship and showed that the social networks of these mitochondria are more than just a handy analogy.
It can be tricky to observe and measure the interactivity of something small in scale and you need to have the right tools. Fortunately, imaging and tracking technologies are getting better all the time.
We use a confocal microscope, and the plant seedlings we use are tiny, around 5mm long.
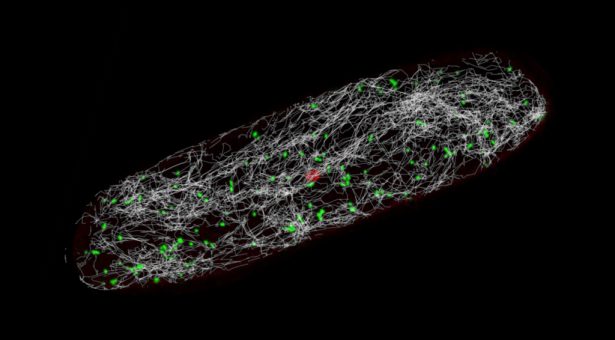
Our plants have a transgene, which localises a green fluorescent protein to the mitochondria, so the individual organelles ‘glow’ brightly when imaged with the correct laser power. This makes them visible in real-time, so you can observe them whizzing about in the cell, going about their very important business of energy provision.
From these videos, we use a tracking software that pinpoints the position of these individual organelles over time, and from this you get a quantification of how many mitochondria there are, where they are at any given time, and how close they are to each other.
We measure interactivity as proximity, so if one mitochondrion is within a close distance to another, we record that.
When you’ve measured all interactions within the video, you can build a social network at any time point in the video.
Each interaction is an edge between the nodes that represent the mitochondria. Borrowing lessons from computer science and graph theory, we can quantify connectivity amongst small groups or cliques and across the whole cell.
The movement of these organelles is dictated by all sorts of cellular signals, as well as the streaming of cytoplasm in the cell, and the crowded cellular environment.
It is hard not to anthropomorphise them, as they often look like people milling around a party – a few who just stick to a certain area, very popular ones, ones who don’t talk to anyone, and one’s who are so busy they just shoot through the room with no time to spare.
We’re trying to find out why these varied behaviours and connectivities might be benefit the cell (and what makes a good party…).
Using social networks to quantify how these mitochondria communicate and connect has allowed us to see what the cell is prioritising.
When you have this system of dynamic organelles, there needs to be a way of viewing connectivity between them all, and social encounter networks have allowed us to do this.
It’s also a tool that can be used across other species, and even other organelles.
We’re currently using it to investigate how mitochondrial movement and connectivity is impacted when the genetics of these individual organelles are perturbed- it’s a direct application of this technique born out of a scoping project- so that’s amazing to see.
This new project started with the curious difference between plant and animal mitochondria.
Animal mitochondria form into long spaghetti-like reticulations, forming a physical conjoined network of organelles.
In plants however they remain mostly punctate, zipping about on their own and only fusing up transiently.
This could be driven by the fact that mitochondria DNA (an heirloom from their bacteria ancestors) recombines in plants, and not in animals.
This partitioning allows the cell to control, through their movement, which mitochondria can speak to which and so have the opportunity to recombine their DNA with.
This recombination also helps plants have very flexible responses to changing environments from which they cannot move away, something animals have less of an issue with.
Our current work delves into this mitochondrial motion in plants, focusing on how it enables this DNA sharing and recombination- and applying our networks to a new problem.
Looking forwards, I’m also really excited for the current research being done on organelle positioning in plant cells, and how this can help us design more efficient connections between organelles such as mitochondria/chloroplasts/peroxisomes and applying that to crop plants and engineered C4 capacities.
In short, mitochondria are amazing as they are so dynamic- they are fascinating to watch and hypothesise about what they are up to- while trying not to anthropomorphise them too much…
I grew up in Worcestershire, in the countryside, and have always loved being outside exploring.
I was always curious, and still am, in everything around me. My favourite things when I was little were treasure hunts, and I got into trouble pulling the keys out of my mum’s early PC keyboard trying to find out what was underneath.
When I first started studying biology I found something that I really loved, for me it was more intuitive than other subjects and I think it was then that being a scientist became a real goal.
My first cell biology lectures at undergraduate level showed me how incredible these biological processes are- for example clathrin coat formation around a vesicle- how can something so tiny be so well orchestrated, so clear in its objective, while also being very real and very measurable?
I was blown away that we can find out so much about this whole new (to me) world – but I didn’t really focus on plants until it came to my undergraduate dissertation.
George Bassel took me on as a summer project student and his team taught me about how we can use computational tools to uncover patterns of connectivity in plants, and how to blend experimental and theoretical data, wet lab work and computational analysis.
I’ve followed that philosophy through to my PhD with Iain Johnston and the Stochastic Biology team, who are now in Bergen (I’m also working with the Gibbs lab here at UoB- keeping me ‘plant-based’), and it’s been a fascinating journey so far- quantitative approaches are the backbone of our work and allow us to ask new questions about a system, testing hypotheses that may otherwise be out of reach.
One thing I think our group does very well (although I am biased) is taking a very broad look at the questions we are addressing- what do these processes look like across the whole tree of life; what can we learn from other disciplines?
I plan to carry that philosophy through to wherever my scientific journey takes me…”

